Data analysis with program PAIREF leads to more accurate structures of biomolecules
Fri Jun 12 13:43:18 CEST 2020
Fri Jun 12 13:43:18 CEST 2020 | Fri Jun 12 13:43:18 CEST 2020 - Fri Jun 12 13:43:18 CEST 2020

Diffraction methods are capable to determine 3D atomic structures of biomolecules (e.g. proteins and DNA), essential for the design of drugs and biotechnological applications, etc. A new tool PAIREF, developed at CTU in close collaboration with IBT, provides a comprehensive analysis of experimental data resolution that leads to an optimal accuracy of the structures.
During the processing of experimental diffraction data, the level separating useful signal from noise (known as the high-resolution limit) has to be estimated. Despite the direct impact of this parameter on the quality of atomic structure, there is no general consensus on criteria to its appropriate determination. Thus, data still containing structural information are often unnecessarily refused or, in contrary, too noisy data are involved in processing and structure determination.
This discrepancy was clarified with the advent of the paired refinement protocol, performing the direct comparison of experimental data and the structure model quality. Program PAIREF, developed in Python, provides full automation of this procedure. It represents a new tool for a more comprehensive analysis of experimental data, innovating the established practise and reviewing the currently used standards, and therefore significant for practical use by macromolecular crystallographers. In our work, we also monitored the impact of the high-resolution limit on the quality of resulting structures. We could observe significant improvements at places in structures difficult to interpret while using the optimal choice of the high-resolution limit, based on the results from the PAIREF program.
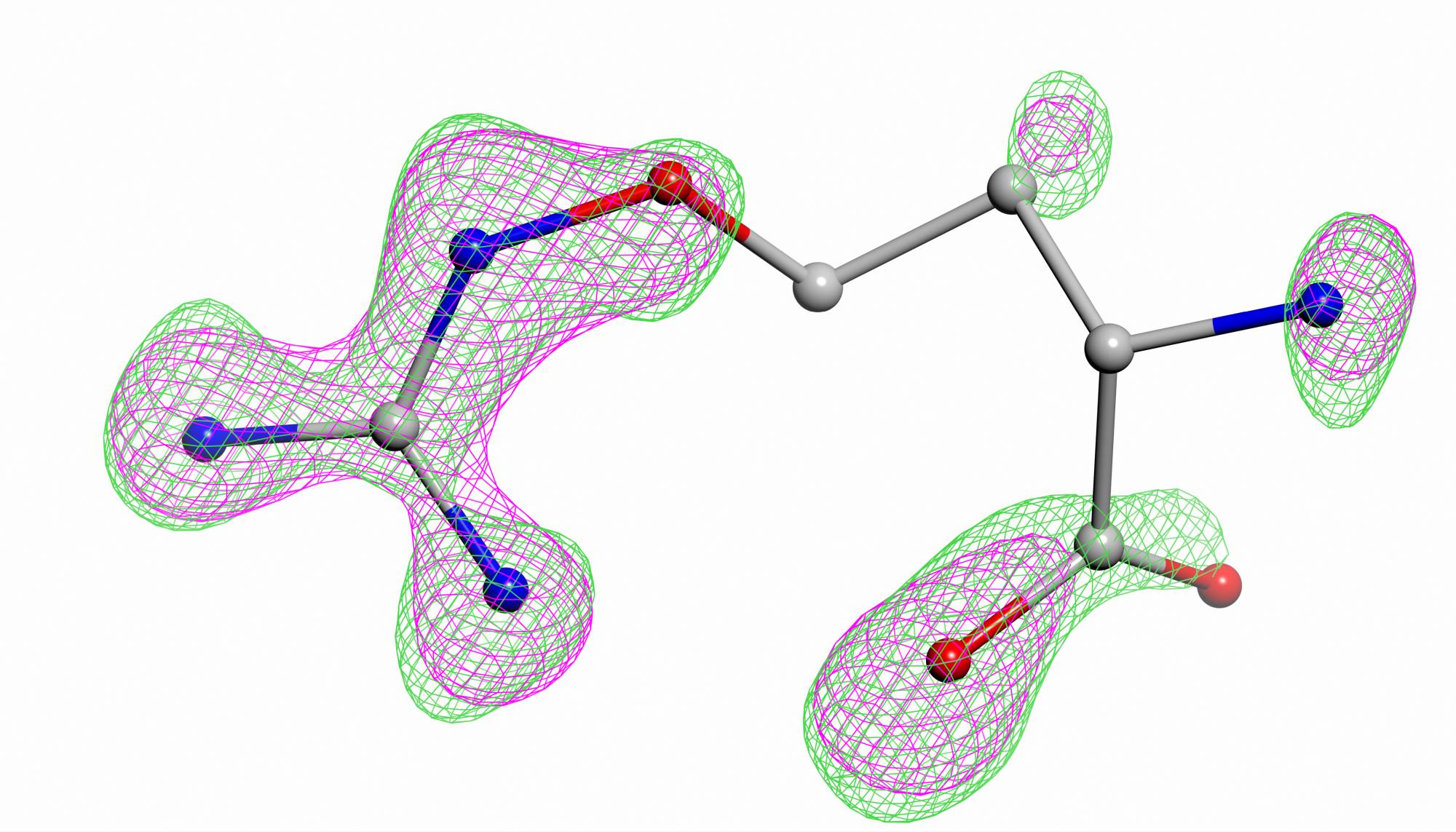
Details about PAIREF developmentand its applicability were published in journal IUCrJ (Malý et al., Paired refinement under the control of PAIREF, IUCrJ 7, 2020, https://doi.org/10.1107/S2052252520005916)).
Project website: https://pairef.fjfi.cvut.cz/
